PCR (Polymerase Chain Reaction)
What is PCR?
A PCR or polymerase chain reaction is a laboratory procedure in which millions of copies of a specific piece of DNA are made. It is essentially an amplification method, whereby the tiniest amounts of DNA that may be present in blood, hair or tissues can be copied so that there is enough for analysis.
The name of this method is derived from the key component involved that carries out the replication of the DNA, called a DNA polymerase. This is an enzyme that exists in nature. The most commonly used polymerase is Taq polymerase, which is obtained from the bacterium Thermus aquaticus. This enzyme works optimally at about 70oC. It can create a new DNA strand, using the original DNA as a template, and using DNA oligonucleotides (also known as primers). The primers used in PCR are synthesized, short sequences of DNA that are made to match exactly the ends of the DNA region to be copied.
PCR has replaced previous methods of DNA replication that used bacteria and could take several weeks to complete. PCR can be done within a few hours, making it a very rapid assay. Speed is often required in a diagnostic setting when urgent results are necessary. PCR was developed around 1983 by Kary Mullis, who won a Nobel Prize in Chemistry for the invention. Since then, PCR has been widely used as a diagnostic and research tool. Its applications are continually growing and are widespread over many scientific disciplines, including molecular biology, microbiology, genetics, clinical diagnostics, forensic science, environmental science, hereditary studies and paternity testing.
Applications of PCR
PCR is extensively used in analysing clinical specimens for the presence of infectious agents, including HIV, hepatitis, human papillomavirus (the causative agent of genital warts and cervical cancer), Epstein-Barr virus (glandular fever), malaria and anthrax.
PCR is particularly invaluable in the early detection of HIV as it can identify the DNA of the virus within human cells immediately following infection, as opposed to the antibodies that are produced weeks or months after infection. PCR can also be used to determine the viral load (i.e. how much virus is circulating around the body), which is a useful measure of prognosis.
Malaria is traditionally diagnosed by identifying malarial parasites (Plasmodium falcipurum) through microscopic analysis of the blood. However, PCR technology has been useful in that it can rapidly identify the species of malaria present. This is important in cases of mixed infection, and also in determining the type of drug treatment to use. Currenty, PCR is used to complement microscopic examination.
PCR can be used to identify the bacterium Bacillus anthracis, the causative agent of anthrax. Because of the need to rapidly diagnose such infections, PCR has become an important tool in detecting the presence of anthrax in clinical specimens. It replaces conventional methods of using a specimen to grow the bacteria in the laboratory, which take at least 24 hours. PCR provides a rapid, sensitive and specific alternative.
The role of PCR in cancer diagnostics
PCR is an invaluable tool as it can provide information on a patient’s prognosis, and predict response or resistance to therapy. Many cancers are characterised by small mutations in certain genes, and this is what PCR is employed to identify.
For example, in acute myeloid leukaemia (AML) the presence of a mutation known as t(8:21) can indicate a good prognosis, as certain drugs are known to be successful in patients who carry this mutation. The presence of a mutation in a gene called Flt3 can identify those patients who are less likely to fully respond to treatment with chemotherapy, and who have a high risk of relapsing following treatment.
PCR can also be applied in monitoring leukaemia patients following treatment, by counting the number of cancerous cells that are still circulating in their bodies.
Genetic diseases and paternity testing
Another important application of PCR is in the analysis of mutations that occur in many genetic diseases (e.g. cystic fibrosis, sickle cell anaemia, phenylketonuria, muscular dystrophy). Because of the sensitivity of PCR, this can be done from a single cell taken from an embryo before birth.
The paternity test is essentially carried out by PCR. A cheek swab is taken from inside the mouth of both parents and the child. The DNA is extracted from the cells obtained and is analysed by PCR. Everyone’s DNA is the same in every cell in the body. A child’s DNA should have part of the mother’s and father’s DNA. Several locations called ‘loci’ on the child’s DNA are examined, and the sequences of these loci are compared to the mother and father to see if there are matches from both parents.
How does PCR work?
There are three basic steps involved in performing a PCR. The steps are repeated 30-40 times in cycles of heating and cooling, with each step taking place at a different temperature.
Components required to carry out a PCR
- A DNA template: The DNA to be copied, usually extracted and purified from blood or other tissue.
- Primers: Single stranded oligonucleotides that match exactly the beginning and end of the DNA template. These are generated synthetically.
- A DNA polymerase (e.g. Taq polymerase): To synthesise the DNA.
- dNTPs (Deoxyribonucleotide triphosphates): The building blocks from which the Taq polymerase can synthesise new DNA. These are added in excess amount.
- A buffer solution: Creates an optimal environment for the reaction to occur in.
- Magnesium chloride salt solution
All of the components are mixed together in one tube in very tiny volumes. The reaction is carried out in an automated machine, known as a thermocycler, which is capable of rapidly increasing and decreasing the temperature.
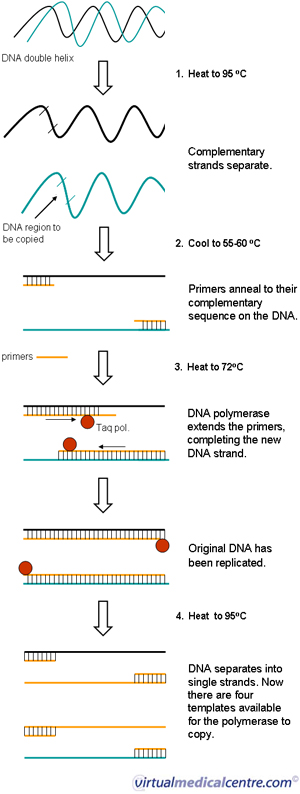
- The first step is known as the denaturation step and is carried out at around 95oC.
DNA exists in nature as a double stranded molecule linked together by weak hydrogen bonds. To be able to copy it, the DNA needs to be separated into single strands (denatured). This can be done by heating it to over 90oC.
- The second step is the annealing step and is carried out at about 55-60oC.
Although the temperature is lowered, an excess amount of primers prevents the denatured DNA from reforming the double helix. The primers attach or ‘anneal’ to their matching sequence on the original DNA strand.
- The final extension step is carried out at 72oC.
Taq DNA polymerase binds to the annealed primer. Taq polymerase works its way along the DNA, adding complementary nucleotides using the dNTPs and other components in the reaction mix. This completes the replication process.
- Once synthesis has been completed, the whole mixture is heated again to 95oC to melt the newly formed DNA complexes. This results in twice the amount of template available for the next round of replication. Repeated heating and cooling quickly amplifies the DNA segment of interest. Roughly one million copies are made after 20 cycles.
Variations of PCR
There are several variations of the PCR technique. One commonly used and important variation is real-time PCR or quantitative PCR (q-PCR). Real-time PCR can be used to count the amount of DNA, or number of copies of a gene, that is present in a sample.
It is employed to determine the viral load of HIV in AIDS patients, and also in cancer diagnostics to count the number of cancerous cells remaining in a patient undergoing treatment.
Limitations and benefits
As with many diagnostic tests in the laboratory, the possibility of false positive and false negative results does exist when PCR is used for detecting infectious agents. Therefore, follow up confirmation tests are always carried out.
References
- Alberts B, Johnson A, Lewis J, Raff M, Roberts K, Walter P. Molecular Biology of the Cell. 4th Ed. New York: Garland Science: Taylor and Francis Group, 2002.
- Mullis KB, Ferre F, Gibbs, RA. The Polymerase chain reaction. Switzerland: Birkhauser Press, 1994.
- Ottesen EA, Hong JW, Quake SR, Leadbetter JR. Microfluidic Digital PCR Enables Multigene Analysis of Individual Environmental Bacteria. 2006; 314(5804): 1464 – 1467
- Prescott LM, Harley JP and Klein DA. Human diseases caused by viruses In: Microbiology. Dubuqu IA: WMC Brown publishers, 1996; p. 729.
- Lodish H et al. The polymerase chain reaction: An alternative to cloning. In: Molecular Cell Biology. New York: Scientific American Books 1998; p. 254.
- Hanscheid T. Diagnosis of malaria: A review of alternatives to conventional microscopy. Clin Lab Haem. 1999: 21: 235-45.
- Hoffmaster AF et al. Evaluation and validation of a real-time polymerase chain reaction assay for rapid identification of Bacillus anthracis. Emerging Infectious Diseases. 2002: 8(10). Available from URL: http://www.cdc.gov/ ncidod/ EID/ vol8no10/ 02-0393.htm
- Bernard PS, Wittwer CT. Real-time PCR technology for cancer diagnostics. Clinical Chemistry. 2002: 48(8): 1178-85.
- Leroy H et al. Prognostic value of real-time quantitative PCR (RQ-PCR) in AML with t (8; 21). Leukaemia. 2005. 19(3): 367-72.
- Abu-Duhier FM et al. FLT3 internal tandem duplication mutations in adult acute myeloid leukaemia define a high-risk group. British Journal of Haematology. 2000; 111(1): 190-5.
- Yang S, Rothman RE. PCR-based diagnostics for infectious diseases: uses, limitations, and future applications in acute-care settings. The Lancet: Infectious diseases. 2004; 4(6); 337-348
- Blanchard MM, Taillon-Miller P, Nowotny P, et al.PCR buffer optimisation with uniform temperature regimen to facilitate automation. Genome Res. 1993; 2: 234-240
- Heid CA, Stevens J, Livak KJ, et al. Real time quantitative PCR. Genome Res.1996 6: 986-994.
- Ray PF, Handyside AH. Increasing the denaturation temperature during the first cycles of amplification reduces allele dropout from single cells for preimplantation genetic diagnosis. Molecular Human Reproduction. 1996; 2(3): 213-218.
Dates
Tags
Created by:
 Login
Login














